calorine: A Python package for constructing and sampling neuroevolution potential models
E. Lindgren,
J. M. Rahm,
E. Fransson,
F. Eriksson,
N. Österbacka,
Z. Fan,
and
P. Erhart
Journal of Open Source Software 9, 6264
(2024)
doi: 10.21105/joss.06264
zenodo: 7919206
(associated data)
Download PDF

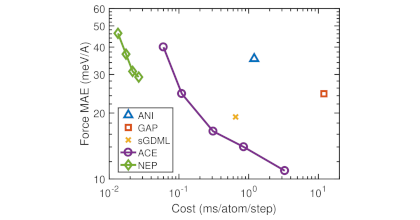
Molecular dynamics (MD) simulations are a key tool in computational chemistry, physics, and materials science, aiding the understanding of microscopic processes but also guiding the development of novel materials. A MD simulation requires a model for the interatomic interactions. To this end, one traditionally often uses empirical interatomic potentials or force fields, which are fast but inaccurate, or ab-initio methods based on electronic structure theory such as density functional theory, which are accurate but computationally very expensive. Machine-learned interatomic potentials (MLIPs) have in recent years emerged as an alternative to these approaches, combining the speed of heuristic force fields with the accuracy of ab-initio techniques. Neuroevolution potentials (NEPs), implemented in the GPUMD package, in particular, are a highly accurate and efficient class of MLIPs. Here, we present calorine, a Python package that simplifies the construction, analysis and use of NEP models via GPUMD. calorine provides a Python interface that makes it easy to access the functionality of GPUMD and integrate it in Python based workflows. This includes but is not limited to managing the construction of NEP models as well as setting up and analyzing MD simulations. The full documentation for calorine in addition to examples and tutorials can be found at https://calorine.materialsmodeling.org/.